Cells have evolved specific machineries in a defined cellular environment to boost metabolic activities. The Liu Lab aims to explore the molecular basis underlying the self assembly and functional regulations of biological machineries. Emerging questions to be addressed include:
(1) How does nature develop functional machineries to enhance cellular metabolism?
(2) How do the structures of biological machineries ensure their physiological functions?
(3) How are the structure and function of the machineries regulated in response to the changing environment?
Advanced knowledge of the biological machinery will underpin the synthetic engineering of new biological “devices” to power cellular metabolism, and provide biotechnological solutions to grand challenges such as global food and energy security. Our research is funded by the UK Royal Society, Biotechnology and Biological Sciences Research Council (BBSRC), Leverhulme Trust, China Scholarship Council, Biochemistry Society, Marie Curie Fellowship.
1. Bacterial microcompartments
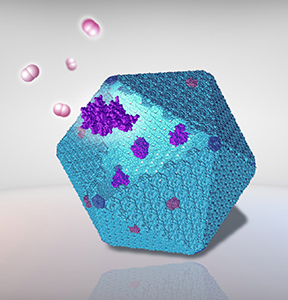
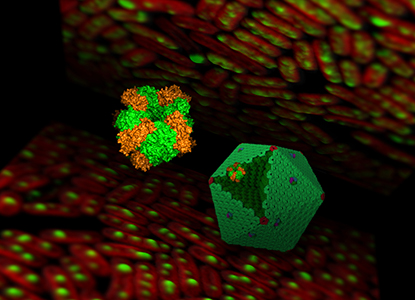
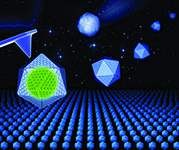
Bacterial microcompartments (BMCs) are proteinaceous organelles widespread among bacterial phyla. They compartmentalize enzymes within a selectively permeable shell and play important roles in carbon fixation, pathogenesis, and microbial ecology. We focus on the biogenesis and self-assembly of carboxysomes from cyanobacteria and propanediol utilization (Pdu) microcompartments from the pathogenic Salmonella enterica, to provide insight into the structural principles of bacterial protein organelles and inform strategies to engineer functional bacterial microcompartments in heterologous hosts for metabolic improvement.
Our main findings involve:
(1) quantifcation of the absolute protein stoichiometry of carboxysomes and Pdu metabolosomes (Nature Communications 2020, Plant Cell 2019).
(2)
characterisation of the roles of auxiliary proteins and environmental factors in mediating carboxysome biogenesis and organisational regulation in bacteria (PNAS 2020, Life 2020, Plant Physiology 2019, Plant Physiology 2016).
(3) visualisation of the self-assembly dynamics and sidedness of building proteins in forming bacterial microcompartment shell facets (Nano Letters 2016), environmental regulations (Nanoscale Research Letters 2019), and the nanomechanical properties of carboxysomes (Nanoscale 2017).
(4)
bioengineering a carboxysome-based nanobioreactor to boost hydrogen production (Nature Commun 2020).
(5) bioengineering of cyanobacterial carboxysomes into the heterologous organism E. coli (Frontiers Plant Sci 2018).
(6) understanding the carboxysome shell permeability using computational simulations (Scientific Reports 2020).
(7)
structural & functional studies on cyanobacterial bicarbonate membrane transporter BicA (Nature Plants 2019).
2. Photosynthetic machinery and membranes
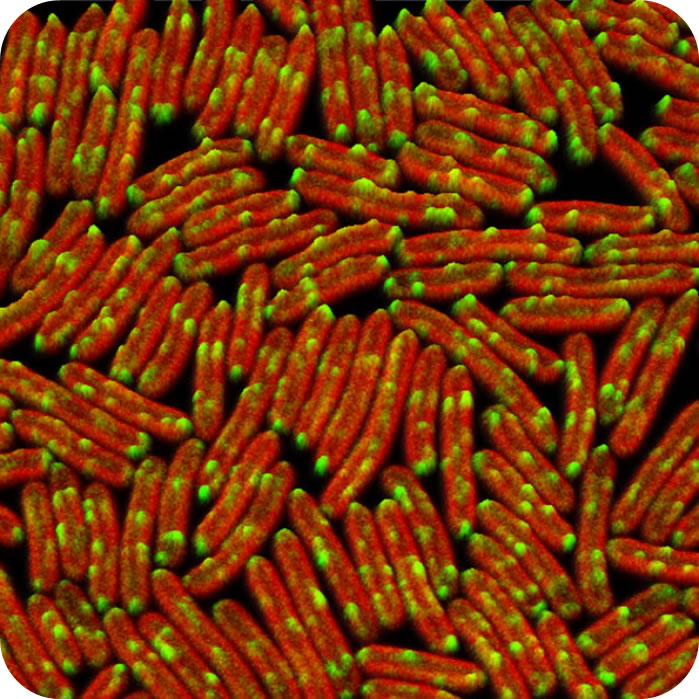
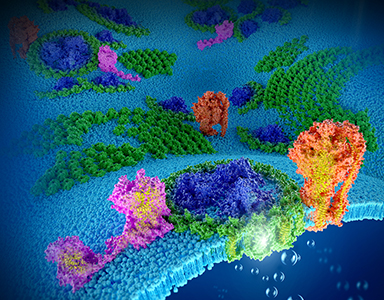
"Without photosynthesis, no complex ecosystems and higher life forms including man would exist." The thylakoid membrane is the site for photosynthetic reactions and responses in cyanobacteria, algae and higher plants [BBA 2016]. We performed direct visualization of the native organization, mobility, and biosynthesis of photosynthetic complexes in photosynthetic membranes from cyanobacteria, purple photosynthetic bacteria and algae. Knowledge and technology obtained from studies on photosynthetic membranes could be extended in general to other biological membranes, such as plant chloroplast and mitochondrial membranes (Annual Review Microbiology 2020, BBA-Bioenergetics 2016, Trends Plant Sci 2013).
Our main findings involve:
(1) deciphering the long-range organisation, local supercomplex organisation, dynamics of photosynthetic complexes in native thylakoid membranes (Nature Plants 2020, Molecular Plant 2017, JBC 2009, PLoS ONE 2009) and mRNA localistion (Nature Plants 2020), using state-of-the-art microscopic techniques including atomic force microscopy, electron microscopy and fluorescence microscopy.
(2) inter- and intra-molecular forces that mediate protein interactions and assembly in photosynthetic machinary (BBA-Bioenergetics 2020, PNAS 2011)
(3)
biogenesis and formation of chromatophores from purple photosynthetic bacteria (BBA-Bioenergetics 2014)
(4) in vivo distribution and dynamics of key respiratory electron donors, type-I NAD(P)H dehydrogenase (NDH-1) and succinate dehydrogenase (SDH) in cyanobacteria (PNAS 2012).